Antibiotic Resistance and Intervention Methods
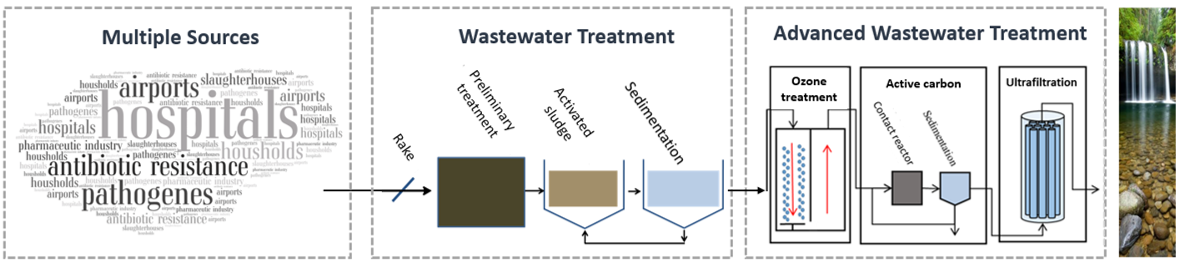
The spread of bacterial pathogens that exhibit insensitivity to common antibiotic therapies is of increasing concern to health authorities worldwide. If this trend continues, there is a risk that the outcome of bacterial infectious diseases will be severe. The situation is also complicated by the fact that research in the pharmaceutical industry for new effective antibiotics is becoming increasingly limited, so that the clinical significance of antibiotic resistance will increase even more in the future. For example, it is predicted that by 2050 infections with antibiotic-resistant and multidrug-resistant bacteria will be responsible for 10 million deaths per year worldwide, replacing cancer as the number one cause of death.
Increasing exposure to clinically-relevant antibiotic-resistant and multidrug-resistant bacteria in the aquatic environment is attributed, among other things, to the discharge of conventionally treated wastewater from sewage treatment plants. The investigation of extended intervention measures to interrupt these dissemination pathways into the environment is a focus of the working group. Modern molecular biological detection systems (qPCR, ddPCR, SeqStudio) are used to detect and quantify facultative pathogenic bacteria of the ESKAPE group and clinically relevant antibiotic resistances, especially against reserve antibiotics, as well as culture-based methods to identify multi-resistant Gram-negative bacteria (ESBL classification). By selecting suitable parameters, this allows both an evaluation of the efficiency of intervention measures to be carried out and dynamics (forecasts/correlations) and biological risk assessments for specific resistance determinants to be carried out. This results in suitable concepts for target-oriented microbiological assessment in order to identify a need for action (see also final report on BMBF HyReKA project).
Prof. Dr. Thomas Schwartz
Microbiology / Molecularbiology
phone:+49721608-26802


